Abstract
Background
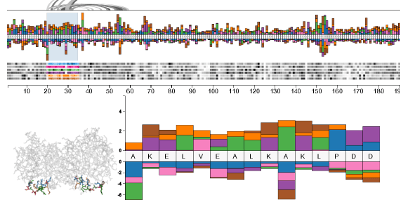
A complete understanding of the relationship between the amino acid sequence and resulting protein function remains an open problem in the biophysical sciences. Current approaches often rely on diagnosing functionally relevant mutations by determining whether an amino acid frequently occurs at a specific position within the protein family. However, these methods do not account for the biophysical properties and the 3D structure of the protein. We have developed an interactive visualization technique, Mu-8, that provides researchers with a holistic view of the differences of a selected protein with respect to a family of homologous proteins. Mu-8 helps to identify areas of the protein that exhibit: (1) significantly different bio-chemical characteristics, (2) relative conservation in the family, and (3) proximity to other regions that have suspect behavior in the folded protein.
Methods
Our approach quantifies and communicates the difference between a reference protein and its family based on amino acid indices or principal components of amino acid index classes, while accounting for conservation, proximity amongst residues, and overall 3D structure.
Results
We demonstrate Mu-8 in a case study with data provided by the 2013 BioVis contest. When comparing the sequence of a dysfunctional protein to its functional family, Mu-8 reveals several candidate regions that may cause function to break down.
Citation
Johnathan D Mercer,
Balaji Pandian,
Alexander Lex,
Nicolas Bonnel,
Hanspeter Pfister
Mu-8: Visualizing Differences between Proteins and their Families
BMC Proceedings, 8(Suppl 2): S5, doi:10.1186/1753-6561-8-S2-S5, 2014.
BibTeX
@article{2014_bmc_mu8,
title = {Mu-8: Visualizing Differences between Proteins and their Families },
author = {Johnathan D Mercer and Balaji Pandian and Alexander Lex and Nicolas Bonnel and Hanspeter Pfister},
journal = {BMC Proceedings},
doi = {10.1186/1753-6561-8-S2-S5},
url = {http://www.biomedcentral.com/1753-6561/8/S2/S5/abstract},
volume = {8},
number = {Suppl 2},
pages = {S5},
month = {aug},
year = {2014}
}
Context
Mu-8 was conceived as a contribution to the 2013 BioVis Contest and was originally developed with students of Harvard’s CS 171 - Visualization class.