Abstract
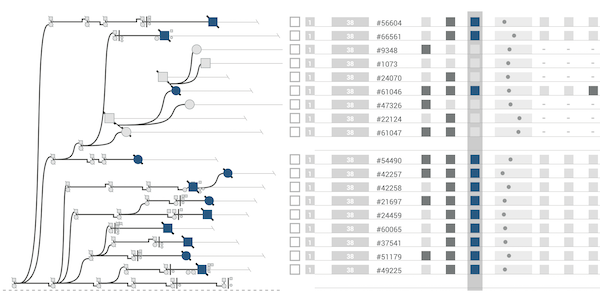
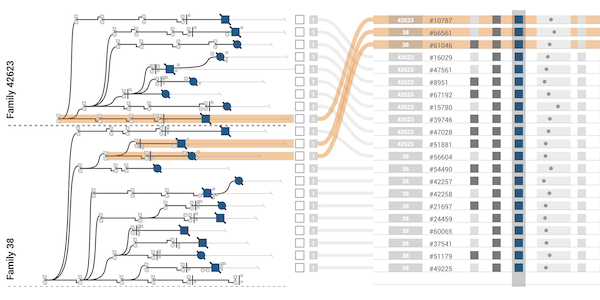
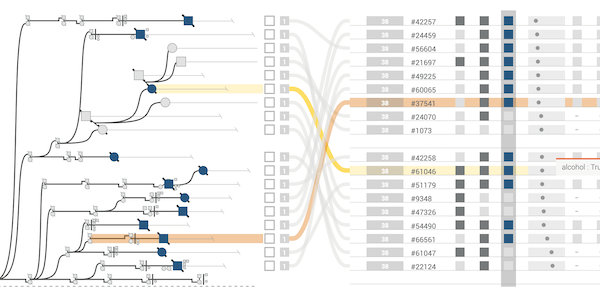
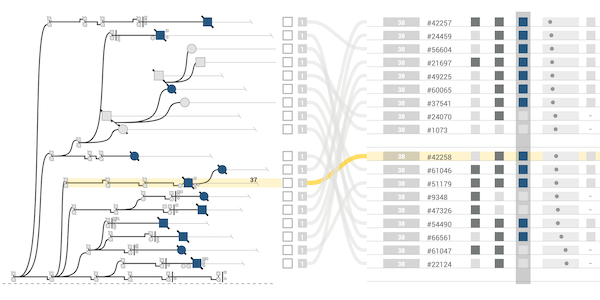
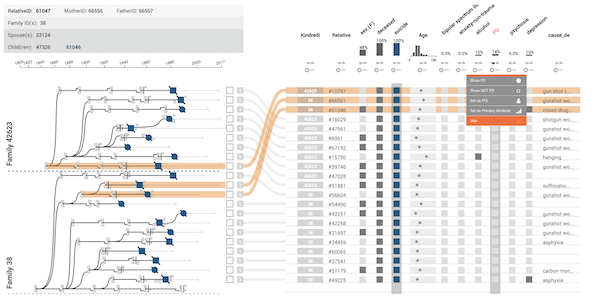
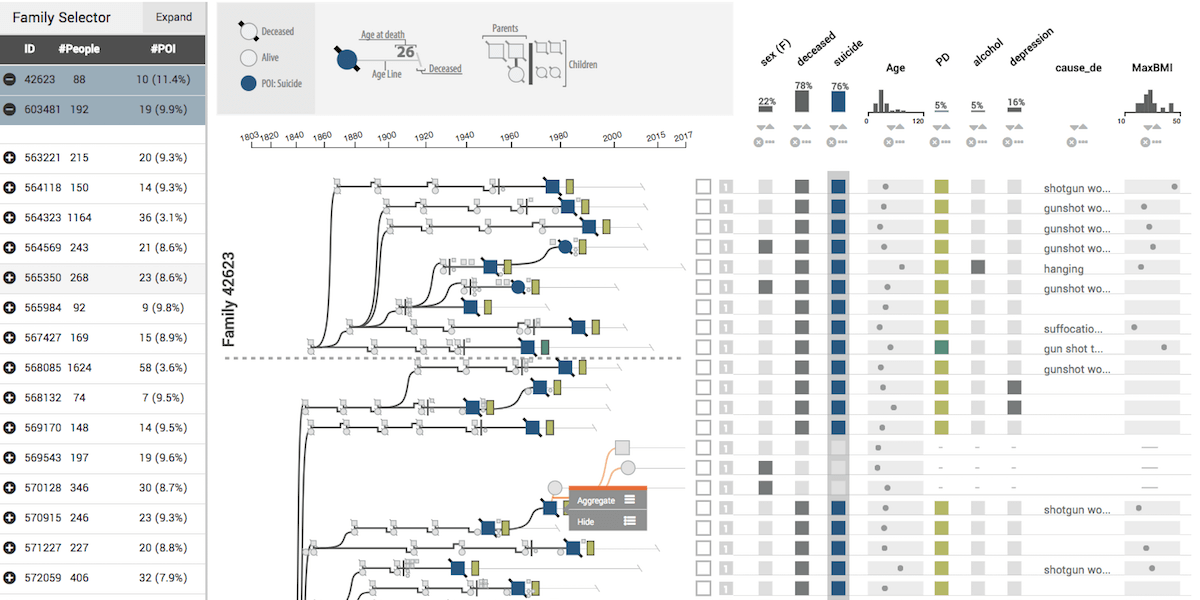
The majority of diseases that are a significant challenge for public and individual heath are caused by a combination of hereditary and environmental factors. In this paper we introduce Lineage, a novel visual analysis tool designed to support domain experts who study such multifactorial diseases in the context of genealogies. Incorporating familial relationships between cases with other data can provide insights into shared genomic variants and shared environmental exposures that may be implicated in such diseases. We introduce a data and task abstraction, and argue that the problem of analyzing such diseases based on genealogical, clinical, and genetic data can be mapped to a multivariate graph visualization problem. The main contribution of our design study is a novel visual representation for tree-like, multivariate graphs, which we apply to genealogies and clinical data about the individuals in these families. We introduce data-driven aggregation methods to scale to multiple families. By designing the genealogy graph layout to align with a tabular view, we are able to incorporate extensive, multivariate attributes in the analysis of the genealogy without cluttering the graph. We validate our designs by conducting case studies with our domain collaborators.Citation
Carolina Nobre,
Nils Gehlenborg,
Hilary Coon,
Alexander Lex
Lineage: Visualizing Multivariate Clinical Data in Genealogy Graphs
IEEE Transactions on Visualization and Computer Graphics, 25(3): 1543-1558, doi:10.1109/TVCG.2018.2811488, 2019.
BibTeX
@article{2018_tvcg_lineage,
title = {Lineage: Visualizing Multivariate Clinical Data in Genealogy Graphs},
author = {Carolina Nobre and Nils Gehlenborg and Hilary Coon and Alexander Lex},
journal = {IEEE Transactions on Visualization and Computer Graphics},
publisher = {IEEE},
doi = {10.1109/TVCG.2018.2811488},
volume = {25},
number = {3},
pages = {1543-1558},
year = {2019}
}
Acknowledgements
We thank Asmaa Aljuhani and Annie Cherkaev for their contributions. We also thank our collaborators and the Visualization Design Lab at the University of Utah for the feedback, and the Caleydo team for their technical support. This work was supported in part by the US National Institutes of Health (U01 CA198935, R00 HG007583, R01MH099134) and the DoD – Office of Economic Adjustment (OEA), ST1605-16-01. We thank the Pedigree and Population Resource of the Huntsman Cancer Institute, University of Utah (funded in part by the Huntsman Cancer Foundation) for its role in the ongoing collection, maintenance, and support of the Utah Population Database (UPDB). We also acknowledge support for the UPDB through grant P30 CA2014 from the National Cancer Institute, and from the University of Utah’s Program in Personalized Health and Center for Clinical and Translational Science.
Images
These images are not part of the original paper and licensed using CC BY 4.0. If you use these images, please cite the paper. Click on the images for full resolution.